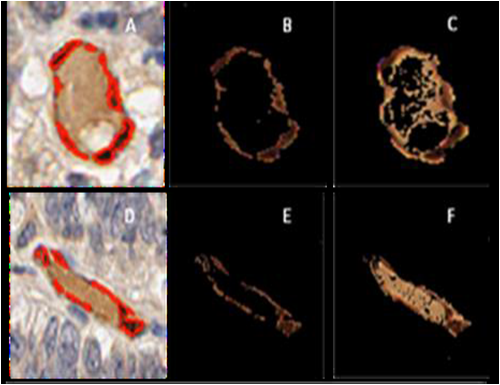
High-throughput Biomarker Quantification
Madabhushi and his team have developed a powerful tool for highthroughput biomarker quantification named Hierarchical Normalized Cuts (HNCuts)15 . The method combines a frequency-weighted version of the mean shift (MS) algorithm with the Normalized Cuts scheme to segment all the image pixels into representative classes. Unlike supervised segmentation strategies, our scheme only requires specification of a small representative sample of the target class to rapidly identify similar objects in the image.
The PIIP will provide software classes needed to transfer information defined by the user within the Sedeen Viewer’s GUI out to the application software. In this case the user will use the viewer’s annotation tools to define the sample color swatch, and this information will be passed out to HNCuts. The PIIP will also allow any parameters needed by the HNCuts algorithm to be exposed to the user to allow for manual tuning. Once processing is carried out, it will be possible to pass segmentation, quantification results back to the Sedeen Viewer for display as a color/contour overlay.